A new method to perform cistrome-wide association studies (Baca et al. pre-print)
Work led by Sylvan Baca on a new method to link genetic determinants of chromatin with complex disease, in collaboration with the Freedman Lab, is now online:
Genetic determinants of chromatin reveal prostate cancer risk mediated by context-dependent gene regulation.
Baca S, Singler C, Zacharia S, Seo J, Morova T, Hach F, Ding Y, Schwarz T, Huang CF, Kalita C, Groha S, Pomerantz MM, Wang V, Linder S, Sweeney CJ, Zwart W, Lack NA, Pasaniuc B, Takeda DY, Gusev A[+], Freedman ML[+]. 2021
This work shows that chromatin and transcription factor activity in prostate tumors is broadly genetically regulated and can be predicted into disease GWAS data to identify regulatory elements associated with prostate cancer risk (which we term a “Cistrome-Wide Association Study” or CWAS). Strikingly, CWAS associated elements are common at loci that lack colocalization with eQTLs, and we hypothesize that CWAS is able to capture genetic effects that are cell-type and context dependent, which would be missed by looking at steady-state gene expression. Also dig into the Methods for a novel approach to infer genetic variants directly from epigenomic data (making this approach applicable to any functional assay without requiring additional genotyping) and to leverage allele-specific variation.
A full analysis pipeline is available.
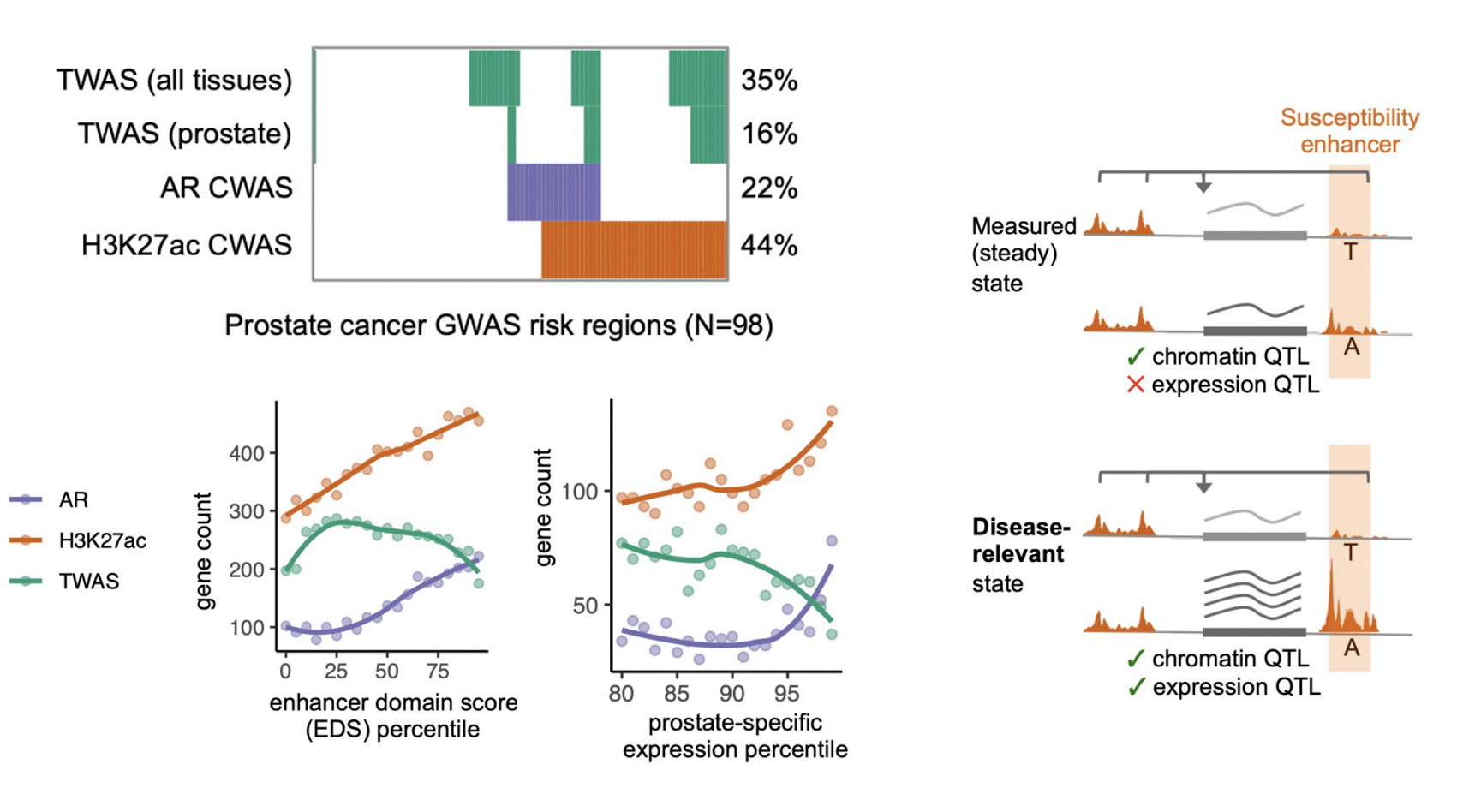
Figure | Summary of CWAS associations (top left); enrichment for context-dependent genes (bottom left); and hypothesized biological model (right).