A new method for genomic classification of cancer of unknown primary and identifying optimal treatments (Moon et al. pre-print)
Work led by Intae Moon on classifying Cancer of Unknown Primary from somatic data is now out as a pre-print
Utilizing Electronic Health Records (EHR) and Tumor Panel Sequencing to Demystify Prognosis of Cancer of Unknown Primary (CUP) patients.
Moon I, LoPiccolo J, Baca SC, Sholl LM, Kehl KL, Hassett MJ, Liu D, Schrag D, Gusev A. 2022
Cancer of Unknown Primary (CUP) is a common cancer type that is defined by having no specified primary tumor after a diagnostic workup. CUP patients typically have dismal outcomes and are challenging to treat because many established therapies are cancer specific. A major open question is whether CUPs are biologically a unique type or a mix of latent, otherwise known cancers. In this work, we develop an algorithm (OncoNPC) that uses somatic mutational data from targeted tumor sequencing to accurately classify cancer types using multi-institutional training data. We apply OncoNPC to a unique cohort of CUP tumor samples with clinical/treatment data to investigate the clinical utility of such an algorithm. OncoNPC CUP subtypes show significant differences in survival outcomes that track with their predicted cancer type and reveal actionable somatic alterations. Integrating germline data, we show that OncoNPC predictions match elevated polygenic risk scores for the predicted cancers – the first indication of a germline genetic correlation between CUP subtypes and known cancers. Finally, in a retrospective causal inference analysis, patients that were treated with therapies that matched the OncoNPC prediction achieved significantly longer survival than those that were discordant. These findings suggest that CUPs exhibit highly distinct molecular subtypes that can be accurately inferred using genomic data and utilized for patient prognosis or to personalize treatments.
For more, see the threads from Intae Moon as well as the related software and data.
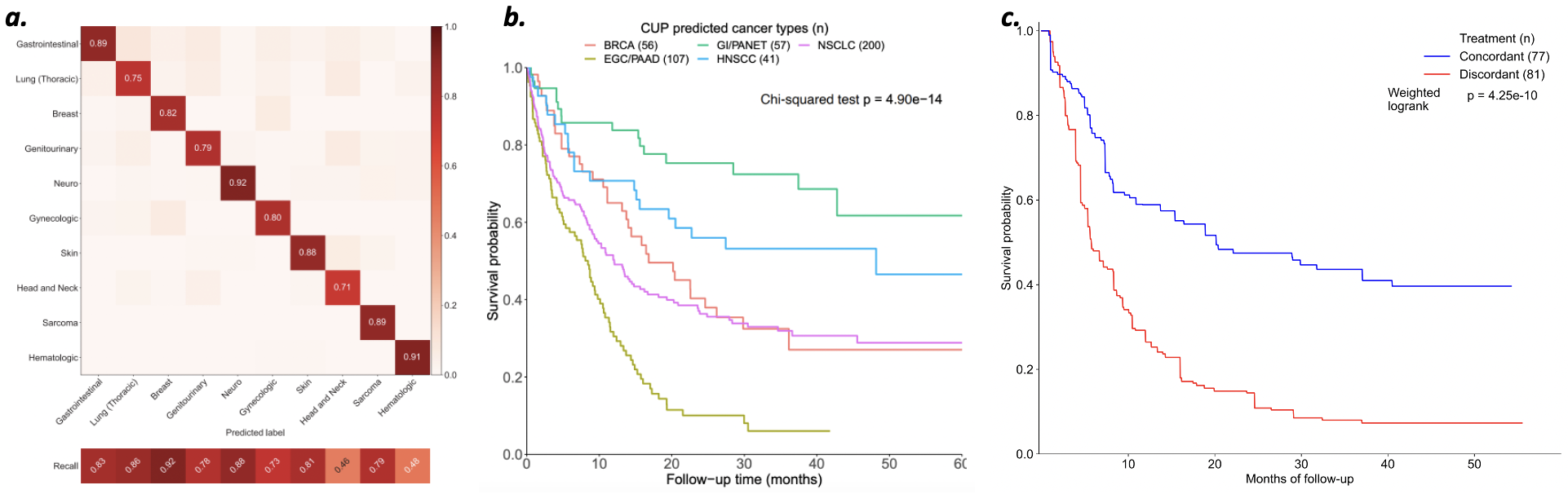
OncoNPC classification accuracy on known tumors (a); OncNPC predicted cancer of unknown primary subtypes show survival differences (b); cancer of unknown primary patients treated consistently with OncoNPC predictions achieved better outcomes (c)