 chromatin TWAS
chromatin TWAS
Supporting Materials for the manuscript Transcriptome-wide association study of schizophrenia and chromatin activity yields mechanistic disease insights. Gusev A, Mancuso N, Won H, Kousi M, Finucane HK, Reshef Y, Song L, Safi A, SCZ Working Group of the Psychiatric Genomics Consortium, McCaroll S, Neale BM, Ophoff RA, O’Donovan MC, Crawford GE, Geschwind DH, Katsanis N, Sullivan PF, Pasaniuc B[+], Price AL[+]. Nature Genetics. 2018
This page provides the documentation and data for analysis of gene expression, chromatin variation, and schizophrenia GWAS. For a general tutorial on TWAS, please see the main documentation at http://gusevlab.org/projects/fusion.
For questions or comments, contact Alexander Gusev [alexander_gusev@dfci.harvard.edu].
SCZ TWAS genes at a glance
Manhattan plot of SCZ TWAS associations from each expression reference panel. Each point represents an individual gene with physical position on the x-axis and (signed) Z-score for association on the y-axis.
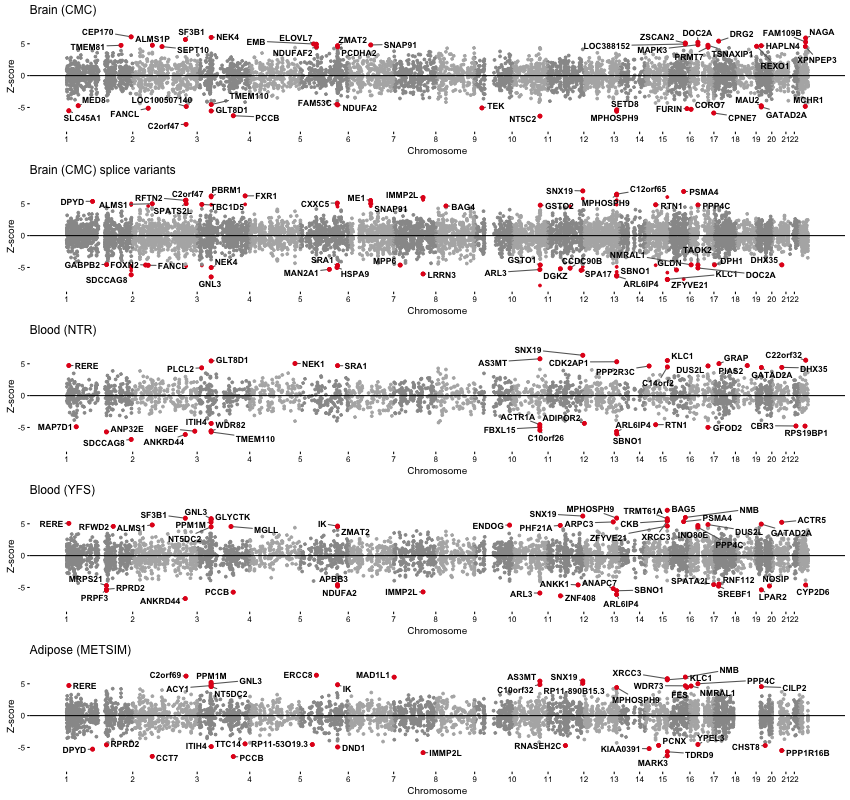
SCZ & chromatin associations in detail
All transcriptome-wide significant TWAS associations and any corresponding chromatin associations.
Study: Expression reference panel used for TWAS (CMC/brain; NTR/blood; YFS/blood; MET/adipose). Z: Z-score for TWAS association, positive sign means increased expression leads to increased risk. INFO: Expected TWAS prediction R2. GWAS P: P-value of the best GWAS SNP in the 1MB locus. joint: Whether this gene was significant in a joint model using all genes (across all expression panels). chromatin: Chromatin TWAS associations (if any). Note: Use search bar to filter chromatin phenotypes (H3k27ac, H3k4me1, H3k4me3, DHS, RPB2, PU1) or reference panels.
Data
| All SCZ TWAS association results across four expression reference panels | [download] |
| All chromatin-TWAS associations | [download] |
| TWAS expression weights | [link] |
| Supplementary Text, Figures, and Tables | [pdf] |
| Associations Spreadsheet | [xlsx] |
Change Log
- 2018/05/09 : Manuscript in print
- 2016/08/02 : Pre-print and data released
- 2016/05/26 : First draft
Logo by Ryan Beck from The Noun Project